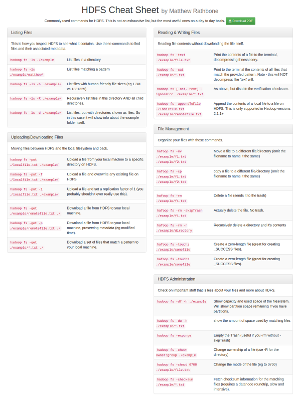
Reading data from HDFS programatically using java (and scala)

Hire me to supercharge your Hadoop and Spark projects
I help businesses improve their return on investment from big data projects. I do everything from software architecture to staff training. Learn More
If you use HDFS, at some point you will need to interact with it programatically. Files can be in several formats, and stored using a spectrum of compression codecs.
Thankfully the HDFS api’s allow you to interact with files fairly easily with only a little boiler plate.
Text Files
In bash you can read any text-format file in hdfs (compressed or not), using the following command:
hadoop fs -text /path/to/your/file.gzIn java or scala you can read a file, or directory of files (taking compression into account) using the function below.
public List<String> readLines(Path location, Configuration conf) throws Exception {
FileSystem fileSystem = FileSystem.get(location.toUri(), conf);
CompressionCodecFactory factory = new CompressionCodecFactory(conf);
FileStatus[] items = fileSystem.listStatus(location);
if (items == null) return new ArrayList<String>();
List<String> results = new ArrayList<String>();
for(FileStatus item: items) {
// ignoring files like _SUCCESS
if(item.getPath().getName().startsWith("_")) {
continue;
}
CompressionCodec codec = factory.getCodec(item.getPath());
InputStream stream = null;
// check if we have a compression codec we need to use
if (codec != null) {
stream = codec.createInputStream(fileSystem.open(item.getPath()));
}
else {
stream = fileSystem.open(item.getPath());
}
StringWriter writer = new StringWriter();
IOUtils.copy(stream, writer, "UTF-8");
String raw = writer.toString();
String[] resulting = raw.split("\n");
for(String str: raw.split("\n")) {
results.add(str);
}
}
return results;
}
// example usage:
Path myfile = new Path("/path/to/results.txt");
List<String> results = readLines(myfile, new Configuration());In fact, this code is taken from my Hadoop test helper library, so I know it works. :-)
Sequence Files
Sequence files are harder to read as you have to read in key/value pairs. Here is a simple function, again taken straight from my Hadoop test helper library.
public <A extends Writable, B extends Writable> List<Tuple<A, B>> readSequenceFile(Path path, Configuration conf, Class<A> acls, Class<B> bcls) throws Exception {
SequenceFile.Reader reader = new SequenceFile.Reader(conf, SequenceFile.Reader.file(path));
long position = reader.getPosition();
A key = acls.newInstance();
B value = bcls.newInstance();
List<Tuple<A, B>> results = new ArrayList<Tuple<A,B>>();
while(reader.next(key,value)) {
results.add(new Tuple(key, value));
key = acls.newInstance();
value = bcls.newInstance();
}
return results;
}
// example usage
List<Tuple<LongWritable, Text>> results = readSequenceFile(new Path("/a/b/c"), new Configuration(), LongWritable.class, Text.class);